Positron Emission Tomography
Support for Positron Emission Tomography (PET) was developed as a BIDS Extension Proposal. Please see Citing BIDS on how to appropriately credit this extension when referring to it in the context of the academic literature.
Example datasets
Several example PET datasets have been formatted using this specification and can be used for practical guidance when curating a new dataset.
Further PET datasets are available from OpenNeuro.
Terminology and conventions
PET-BIDS is fully consistent with the BIDS specification as a whole. However, BIDS was initially developed in the context of MRI, so some terminology may be unfamiliar to researchers from each field. This section adds clarifications to Common Principles - Definitions for the PET context, and introduces the term "time zero" which is currently specific to PET.
-
Session - In most cases, a new session with respect to PET corresponds to a visit to the scanning site, and starts with a new injection. In situations where different data types are obtained over several visits (for example, FDG PET on one day followed by amyloid PET a couple days after) these scans may be grouped into the same session. In other datasets, a subject leaving the scanner and returning under the same injection may be considered separate sessions.
-
Run - In PET, subjects may have to leave the scanner to use the bathroom. While leaving the scanner would interrupt an MR acquisition, in PET this disruption is more appropriately considered missing data during a run.
-
Time zero - A reference point in time, to which all timestamps pertaining to a recording are relative. Time zero will most commonly be the time of injection of a radioisotope, or the time at which the first scan of an acquisition is begun. If a pharmacological within-scan challenge is performed, another time zero may be more convenient.
An overview of a common PET experiment (with blood data) can be seen in Figure 1, defined on a single time scale relative to a predefined "time zero".
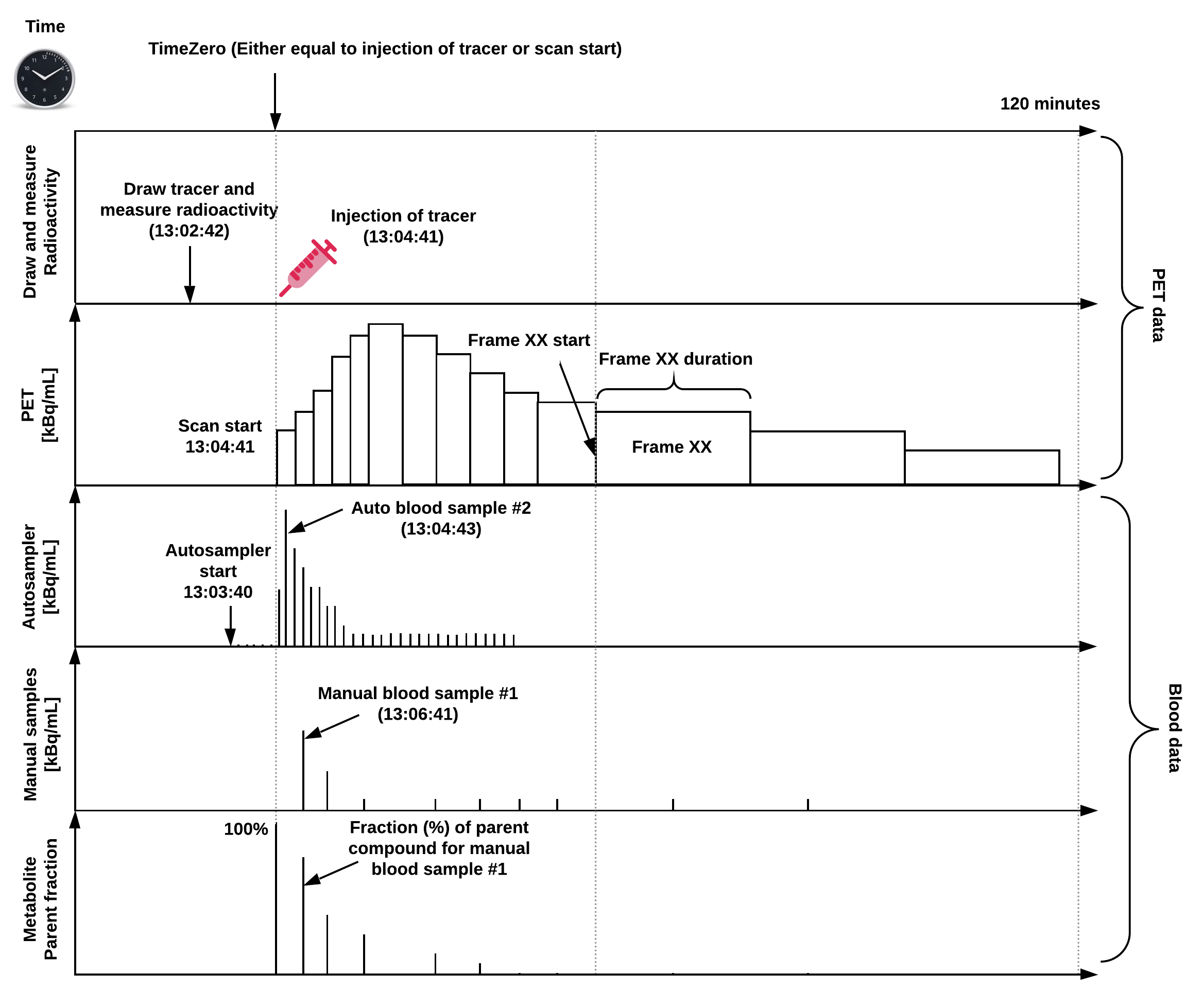
Figure 1: Overview of a common PET experiment, including blood measurements,
and defined on a common time scale.
Note, "time zero" is often defined as time of injection or scan start,
but if a pharmaceutical challenge is carried out during the scan,
this time point may also be chosen as time zero.
The injected dose, the PET data, and blood data should all be decay-corrected to time zero,
but because the time of injection does not always coincide with scan start,
the PET data may not always be decay-corrected to the time of injection.
If this is not the case, this may be indicated in the reconstruction section
(ImageDecayCorrected and ImageDecayCorrectionTime).
In this example, tracer injection coincides with scan start.
PET recording data
Template:
sub-<label>/
[ses-<label>/]
pet/
sub-<label>[_ses-<label>][_task-<label>][_trc-<label>][_rec-<label>][_run-<index>]_pet.json
sub-<label>[_ses-<label>][_task-<label>][_trc-<label>][_rec-<label>][_run-<index>]_pet.nii[.gz]
sub-<label>[_ses-<label>]_task-<label>[_trc-<label>][_rec-<label>][_run-<index>]_events.json
sub-<label>[_ses-<label>]_task-<label>[_trc-<label>][_rec-<label>][_run-<index>]_events.tsv
sub-<label>[_ses-<label>]_task-<label>[_trc-<label>][_rec-<label>][_run-<index>][_recording-<label>]_physio.json
sub-<label>[_ses-<label>]_task-<label>[_trc-<label>][_rec-<label>][_run-<index>][_recording-<label>]_physio.tsv.gz
sub-<label>[_ses-<label>]_task-<label>[_trc-<label>][_rec-<label>][_run-<index>][_recording-<label>]_stim.json
sub-<label>[_ses-<label>]_task-<label>[_trc-<label>][_rec-<label>][_run-<index>][_recording-<label>]_stim.tsv.gz
Legend:
-
For more information about filename elements (for example, entities, suffixes, extensions), follow the links embedded in the filename template.
-
Filename entities or directories between square brackets (for example,
[_ses-<label>]) are OPTIONAL. -
Some entities may only allow specific values, in which case those values are listed in
<>, separated by|. -
_<suffix>means that there are several (>6) valid suffixes for this filename pattern. -
.<extension>means that there are several (>6) valid extensions for this file type. -
[.gz]means that both the unzipped and gzipped versions of the extension are valid.
PET data MUST be stored in the pet directory.
PET imaging data SHOULD be stored in 4D (or 3D, if only one volume was acquired)
NIfTI files with the _pet suffix.
Volumes MUST be stored in chronological order (the order they were acquired in).
The OPTIONAL task-<label> is used to
indicate a task subjects were asked to perform in the scanner.
Those labels MUST be consistent across subjects and sessions.
For task based PET, a corresponding task events file MUST be provided
(please note that this file is not necessary for resting scans).
The trc-<label> entity is used to
indicate the tracer used.
This entity is OPTIONAL if only one tracer is used in the study,
but REQUIRED to distinguish between tracers if multiple are used.
The label used is arbitrary and each file requires a separate JSON sidecar
with details of the tracer used (see below).
Examples are trc-18FFDG for fludeoxyglucose or trc-11CPIB for Pittsburgh compound B.
Other labels are permitted, as long as they are consistent across subjects and sessions
and consist only of the legal label characters.
If more than one run of the same task and acquisition (tracer) are acquired during
the same session, the run-<index> entity MUST be used:
_run-1, _run-2, _run-3, and so on.
If only one run was acquired the run-<index> can be omitted.
The OPTIONAL rec-<label> entity
is used to indicate the reconstruction method used for the image,
with four reserved values:
acdyn, for reconstructions with attenuation correction of dynamic data;acstat, for reconstructions with attenuation correction of static data;nacdyn, for reconstructions without attenuation correction of dynamic data;nacstat, for reconstructions without attenuation correction of static data.
Further details regarding reconstruction are in the _pet.json file.
If multiple reconstructions of the data are made with the same type of reconstruction,
a number MAY be appended to the label,
for example rec-acdyn1 and rec-acdyn2.
Shared MRI data along with PET
PET and MRI images may be aggregated in the same dataset.
When analyzing MRI and PET data together,
it is essential to specify whether MR images have been corrected for gradient non-linearities,
using the NonLinearGradientCorrection metadata field
(see Sequence Specifics),
which is REQUIRED for all MR data if PET data is also present in the dataset
(see also PET-MRI correspondence).
In the case of studies using combined PET/fMRI,
subject-specific tasks may be carried out during the acquisition within the same session.
If the same task is recorded with both modalities,
the same task-<label> entity SHOULD be used.
For further details, see
Task (including resting state) imaging data.
In addition to the imaging data (*.nii) a _pet.json sidecar file MUST be provided.
The included metadata are divided into sections described below.
PET metadata
PET data MUST be described by metadata fields, stored in sidecar JSON files. These fields are derived from the recommendations in Knudsen et al. 2020, doi:10.1177/0271678X20905433, which we divide into several categories:
Radiochemistry
| Key name | Requirement Level | Data type | Description |
|---|---|---|---|
| TracerName | REQUIRED | string | Name of the tracer compound used (for example, "CIMBI-36") Corresponds to DICOM Tags (0008,0105) Mapping Resource and (0008,0122) Mapping Resource Name. |
| TracerRadionuclide | REQUIRED | string | Radioisotope labeling tracer (for example, "C11"). Corresponds to DICOM Tags (0008,0104) CodeValue and (0008,0104) CodeMeaning. |
| InjectedRadioactivity | REQUIRED | number | Total amount of radioactivity injected into the patient (for example, 400). For bolus-infusion experiments, this value should be the sum of all injected radioactivity originating from both bolus and infusion. Corresponds to DICOM Tag 0018, 1074 Radionuclide Total Dose. |
| InjectedRadioactivityUnits | REQUIRED | string | Unit format of the specified injected radioactivity (for example, "MBq"). |
| InjectedMass | REQUIRED | number or "n/a" |
Total mass of radiolabeled compound injected into subject (for example, 10). This can be derived as the ratio of the "InjectedRadioactivity" and "MolarRadioactivity". For those tracers in which injected mass is not available (for example FDG) can be set to "n/a"). |
| InjectedMassUnits | REQUIRED | string or "n/a" |
Unit format of the mass of compound injected (for example, "ug" or "umol"). Note this is not REQUIRED for an FDG acquisition, since it is not available, and SHOULD be set to "n/a". |
| SpecificRadioactivity | REQUIRED | number or "n/a" |
Specific activity of compound injected. Note this is not REQUIRED for an FDG acquisition, since it is not available, and SHOULD be set to "n/a". |
| SpecificRadioactivityUnits | REQUIRED | string or "n/a" |
Unit format of specified specific radioactivity (for example, "Bq/g"). Note this is not REQUIRED for an FDG acquisition, since it is not available, and SHOULD be set to "n/a". |
| ModeOfAdministration | REQUIRED | string | Mode of administration of the injection (for example, "bolus", "infusion", or "bolus-infusion"). |
| TracerRadLex | RECOMMENDED | string | ID of the tracer compound from the RadLex Ontology. |
| TracerSNOMED | RECOMMENDED | string | ID of the tracer compound from the SNOMED Ontology (subclass of Radioactive isotope). |
| TracerMolecularWeight | RECOMMENDED | number | Accurate molecular weight of the tracer used. |
| TracerMolecularWeightUnits | RECOMMENDED | string | Unit of the molecular weights measurement (for example, "g/mol"). |
| InjectedMassPerWeight | RECOMMENDED | number | Injected mass per kilogram bodyweight. |
| InjectedMassPerWeightUnits | RECOMMENDED | string | Unit format of the injected mass per kilogram bodyweight (for example, "ug/kg"). |
| SpecificRadioactivityMeasTime | RECOMMENDED | string | Time to which specific radioactivity measurement above applies in the default unit "hh:mm:ss". |
| MolarActivity | RECOMMENDED | number | Molar activity of compound injected. Corresponds to DICOM Tag 0018, 1077 Radiopharmaceutical Specific Activity. |
| MolarActivityUnits | RECOMMENDED | string | Unit of the specified molar radioactivity (for example, "GBq/umol"). |
| MolarActivityMeasTime | RECOMMENDED | string | Time to which molar radioactivity measurement above applies in the default unit "hh:mm:ss". |
| InfusionRadioactivity | RECOMMENDED, but REQUIRED if ModeOfAdministration is 'bolus-infusion' |
number | Amount of radioactivity infused into the patient. This value must be less than or equal to the total injected radioactivity ("InjectedRadioactivity"). Units should be the same as "InjectedRadioactivityUnits". |
| InfusionStart | RECOMMENDED, but REQUIRED if ModeOfAdministration is 'bolus-infusion' |
number | Time of start of infusion with respect to "TimeZero" in the default unit seconds. |
| InfusionSpeed | RECOMMENDED, but REQUIRED if ModeOfAdministration is 'bolus-infusion' |
number | If given, infusion speed. |
| InfusionSpeedUnits | RECOMMENDED, but REQUIRED if ModeOfAdministration is 'bolus-infusion' |
string | Unit of infusion speed (for example, "mL/s"). |
| InjectedVolume | RECOMMENDED, but REQUIRED if ModeOfAdministration is 'bolus-infusion' |
number | Injected volume of the radiotracer in the unit "mL". |
| Purity | RECOMMENDED | number | Purity of the radiolabeled compound (between 0 and 100%). Must be a number greater than or equal to 0 and less than or equal to 100. |
Pharmaceuticals
| Key name | Requirement Level | Data type | Description |
|---|---|---|---|
| PharmaceuticalName | RECOMMENDED | string | Name of pharmaceutical coadministered with tracer. Corresponds to DICOM Tag 0008, 0034 Intervention Drug Name. |
| PharmaceuticalDoseAmount | RECOMMENDED | number or array of numbers | Dose amount of pharmaceutical coadministered with tracer. Corresponds to DICOM Tag 0008, 0028 Intervention Drug Dose. |
| PharmaceuticalDoseUnits | RECOMMENDED | string | Unit format relating to pharmaceutical dose (for example, "mg" or "mg/kg"). |
| PharmaceuticalDoseRegimen | RECOMMENDED | string | Details of the pharmaceutical dose regimen. Either adequate description or short-code relating to regimen documented elsewhere (for example, "single oral bolus"). |
| PharmaceuticalDoseTime | RECOMMENDED | number or array of numbers | Time of administration of pharmaceutical dose, relative to time zero. For an infusion, this should be a vector with two elements specifying the start and end of the infusion period. For more complex dose regimens, the regimen description should be complete enough to enable unambiguous interpretation of "PharmaceuticalDoseTime". Unit format of the specified pharmaceutical dose time MUST be seconds. Corresponds to a combination of DICOM Tags (0008,0027) Intervention Drug Stop Time and (0008,0035) Intervention Drug Start Time. |
| Anaesthesia | OPTIONAL | string | Details of anaesthesia used, if any. |
Time
| Key name | Requirement Level | Data type | Description |
|---|---|---|---|
| TimeZero | REQUIRED | string | Time zero to which all scan and/or blood measurements have been adjusted to, in the unit "hh:mm:ss". This should be equal to "InjectionStart" or "ScanStart". |
| ScanStart | REQUIRED | number | Time of start of scan with respect to TimeZero in the default unit seconds. |
| InjectionStart | REQUIRED | number | Time of start of injection with respect to "TimeZero" in the default unit seconds. This corresponds to DICOM Tag 0018, 1072 Contrast/Bolus Start Time converted to seconds relative to "TimeZero". Corresponds to DICOM Tag 0018, 1072 Radiopharmaceutical Start Time. |
| FrameTimesStart | REQUIRED | array of numbers | Start times for all frames relative to "TimeZero" in default unit seconds. |
| FrameDuration | REQUIRED | array of numbers | Time duration of each frame in default unit seconds. This corresponds to DICOM Tag 0018, 1242 Actual Frame Duration converted to seconds. |
| InjectionEnd | RECOMMENDED | number | Time of end of injection with respect to "TimeZero" in the default unit seconds. Corresponds to DICOM Tag 0018, 1073 Radiopharmaceutical Stop Time converted to seconds relative to TimeZero. |
| ScanDate | DEPRECATED | string | Date of scan in the format "YYYY-MM-DD[Z]". This field is DEPRECATED, and this metadata SHOULD be recorded in the acq_time column of the corresponding Scans file. Corresponds to DICOM Tag 0008, 0022 Acquisition Date. |
We refer to the common principles for the standards for describing dates and timestamps, including possibilities for deidentification (see Units).
Reconstruction
| Key name | Requirement Level | Data type | Description |
|---|---|---|---|
| AcquisitionMode | REQUIRED | string | Type of acquisition of the PET data (for example, "list mode"). |
| ImageDecayCorrected | REQUIRED | boolean | Boolean flag specifying whether the image data have been decay-corrected. Must be one of: "true", "false". |
| ImageDecayCorrectionTime | REQUIRED | number | Point in time from which the decay correction was applied with respect to "TimeZero" in the default unit seconds. |
| ReconMethodName | REQUIRED | string | Reconstruction method or algorithm (for example, "3d-op-osem"). This partly matches the DICOM Tag 0054, 1103 Reconstruction Method. |
| ReconMethodParameterLabels | REQUIRED | array of strings | Names of reconstruction parameters (for example, ["subsets", "iterations"]). This partly matches the DICOM Tag 0054, 1103 Reconstruction Method. |
| ReconMethodParameterUnits | RECOMMENDED, but REQUIRED if ReconMethodParameterLabels does not contain "none" |
array of strings | Unit of reconstruction parameters (for example, ["none", "none"]). This partly matches the DICOM Tag 0054, 1103 Reconstruction Method. |
| ReconMethodParameterValues | RECOMMENDED, but REQUIRED if ReconMethodParameterLabels does not contain "none" |
array of numbers | Values of reconstruction parameters (for example, [21, 3]). This partly matches the DICOM Tag 0054, 1103 Reconstruction Method. |
| ReconFilterType | REQUIRED | string or array of strings | Type of post-recon smoothing (for example, ["Shepp"]). This partly matches the DICOM Tag 0018, 1210 Convolution Kernel. |
| ReconFilterSize | RECOMMENDED, but REQUIRED if ReconFilterType is not "none" |
number or array of numbers | Kernel size of post-recon filter (FWHM) in default units "mm". This partly matches the DICOM Tag 0018, 1210 Convolution Kernel. |
| AttenuationCorrection | REQUIRED | string | Short description of the attenuation correction method used. This corresponds to DICOM Tag 0054, 1101 Attenuation Correction Method. |
| ReconMethodImplementationVersion | RECOMMENDED | string | Identification for the software used, such as name and version. |
| AttenuationCorrectionMethodReference | RECOMMENDED | string | Reference paper for the attenuation correction method used. |
| ScaleFactor | RECOMMENDED | array of numbers | Scale factor for each frame. This field MUST be defined if the imaging data (.nii[.gz]) are scaled. If this field is not defined, then it is assumed that the scaling factor is 1. Defining this field when the scaling factor is 1 is RECOMMENDED, for the sake of clarity. |
| ScatterFraction | RECOMMENDED | array of numbers | Scatter fraction for each frame (Units: 0-100%). Corresponds to DICOM Tag 0054, 1323 Scatter Fraction Factor. |
| DecayCorrectionFactor | RECOMMENDED | array of numbers | Decay correction factor for each frame. Corresponds to DICOM Tag 0054, 1321 Decay Factor. |
| DoseCalibrationFactor | RECOMMENDED | number | Multiplication factor used to transform raw data (in counts/sec) to meaningful unit (Bq/ml). Corresponds to DICOM Tag 0054, 1322 Dose Calibration Factor. Corresponds to DICOM Tag 0054, 1322 Dose Calibration Factor. |
| PromptRate | RECOMMENDED | array of numbers | Prompt rate for each frame (same units as Units, for example, "Bq/mL"). |
| SinglesRate | RECOMMENDED | array of numbers | Singles rate for each frame (same units as Units, for example, "Bq/mL"). |
| RandomRate | RECOMMENDED | array of numbers | Random rate for each frame (same units as "Units", for example, "Bq/mL"). |
All reconstruction-specific parameters that are not specified, but one wants to include, should go into the ReconMethodParameterValues field.
Hardware information
| Key name | Requirement Level | Data type | Description |
|---|---|---|---|
| Manufacturer | REQUIRED | string | Manufacturer of the equipment that produced the measurements. Corresponds to DICOM Tag 0008, 0070 Manufacturer. |
| ManufacturersModelName | REQUIRED | string | Manufacturer's model name of the equipment that produced the measurements. Corresponds to DICOM Tag 0008, 1090 Manufacturers Model Name. |
| Units | REQUIRED | string | Measurement units for the associated file. SI units in CMIXF formatting are RECOMMENDED (see Units). SI unit for radioactivity (Becquerel) should be used (for example, "Bq/mL"). Corresponds to DICOM Tag 0054, 1001 Units. |
| BodyPart | RECOMMENDED | string | Body part of the organ / body region scanned. Corresponds to DICOM Tag 0018, 0015 Body Part Examined. |
Institution information
| Key name | Requirement Level | Data type | Description |
|---|---|---|---|
| InstitutionName | RECOMMENDED | string | The name of the institution in charge of the equipment that produced the measurements. Corresponds to DICOM Tag 0008, 0080 InstitutionName. |
| InstitutionAddress | RECOMMENDED | string | The address of the institution in charge of the equipment that produced the measurements. Corresponds to DICOM Tag 0008, 0081 InstitutionAddress. |
| InstitutionalDepartmentName | RECOMMENDED | string | The department in the institution in charge of the equipment that produced the measurements. Corresponds to DICOM Tag 0008, 1040 Institutional Department Name. |
Tissue description
| Key name | Requirement Level | Data type | Description |
|---|---|---|---|
| BodyPart | OPTIONAL | string | Body part of the organ / body region scanned. Corresponds to DICOM Tag 0018, 0015 Body Part Examined. |
| BodyPartDetails | OPTIONAL | string | Additional details about body part or location (for example: "corpus callosum"). |
| BodyPartDetailsOntology | OPTIONAL | string | URI of ontology used for BodyPartDetails (for example: "https://www.ebi.ac.uk/ols/ontologies/uberon"). |
Task
If the OPTIONAL task-<label> is used,
the following metadata SHOULD be used.
| Key name | Requirement Level | Data type | Description |
|---|---|---|---|
| TaskName | RECOMMENDED | string | Name of the task. No two tasks should have the same name. The task label included in the filename is derived from this "TaskName" field by removing all non-alphanumeric characters (that is, all except those matching [0-9a-zA-Z]). For example "TaskName" "faces n-back" or "head nodding" will correspond to task labels facesnback and headnodding, respectively. If used to denote resting scans, a RECOMMENDED convention is to use labels beginning with rest. |
| Instructions | RECOMMENDED | string | Text of the instructions given to participants before the recording. This is especially important in context of resting state recordings and distinguishing between eyes open and eyes closed paradigms. |
| TaskDescription | RECOMMENDED | string | Longer description of the task. |
| CogAtlasID | RECOMMENDED | string | URI of the corresponding Cognitive Atlas Task term. |
| CogPOID | RECOMMENDED | string | URI of the corresponding CogPO term. |
Example (*_pet.json)
{
"Manufacturer": "Siemens",
"ManufacturersModelName": "High-Resolution Research Tomograph (HRRT, CTI/Siemens)",
"BodyPart": "Brain",
"Units": "Bq/mL",
"TracerName": "CIMBI-36",
"TracerRadionuclide": "C11",
"TracerMolecularWeight": 380.28,
"TracerMolecularWeightUnits": "g/mol",
"InjectedRadioactivity": 573,
"InjectedRadioActivityUnits": "MBq",
"InjectedMass": 0.62,
"InjectedMassUnits": "ug",
"SpecificRadioactivity": 929.6,
"SpecificRadioactivityUnits": "MBq/ug",
"ModeOfAdministration": "bolus",
"MolarActivity": 353.51,
"MolarActivityUnits": "GBq/umol",
"MolarActivityMeasTime": "13:04:42",
"TimeZero": "13:04:42",
"ScanStart": 0,
"InjectionStart": 0,
"FrameTimesStart": [0, 10, 20, 30, 40, 50, 60, 80, 100, 120, 140, 160, 180, 240, 300, 360, 420, 480, 540, 660, 780, 900, 1020, 1140, 1260, 1380, 1500, 1800, 2100, 2400, 2700, 3000, 3300, 3600, 3900, 4200, 4500, 4800, 5100, 5400, 5700, 6000, 6300, 6600, 6900],
"FrameDuration": [10, 10, 10, 10, 10, 10, 20, 20, 20, 20, 20, 20, 60, 60, 60, 60, 60, 60, 120, 120, 120, 120, 120, 120, 120, 120, 300, 300, 300, 300, 300, 300, 300, 300, 300, 300, 300, 300, 300, 300, 300, 300, 300, 300, 300],
"AcquisitionMode": "list mode",
"ImageDecayCorrected": true,
"ImageDecayCorrectionTime": 0,
"ReconMethodName": "3D-OSEM-PSF",
"ReconMethodParameterLabels": ["subsets","iterations"],
"ReconMethodParameterUnits": ["none","none"],
"ReconMethodParameterValues": [16,10],
"ReconFilterType": "none",
"AttenuationCorrection": "[137Cs]transmission scan-based"
}
Recommended patient data
Knudsen et al. 2020 (doi:10.1177/0271678X20905433) recommends recording participant body weight. If recorded once per participant, these data SHOULD be included in the Participants file or as Phenotypic and assessment data.
For example:
participant_id body_weight
sub-01 58
sub-02 96
sub-03 72
If multiple measurements are made, these data SHOULD be included in the Sessions file.
For example:
session_id body_weight
ses-01 58
ses-02 59
Blood recording data
Template:
sub-<label>/
[ses-<label>/]
pet/
sub-<label>[_ses-<label>][_task-<label>][_trc-<label>][_rec-<label>][_run-<index>]_recording-<label>_blood.json
sub-<label>[_ses-<label>][_task-<label>][_trc-<label>][_rec-<label>][_run-<index>]_recording-<label>_blood.tsv
Legend:
-
For more information about filename elements (for example, entities, suffixes, extensions), follow the links embedded in the filename template.
-
Filename entities or directories between square brackets (for example,
[_ses-<label>]) are OPTIONAL. -
Some entities may only allow specific values, in which case those values are listed in
<>, separated by|. -
_<suffix>means that there are several (>6) valid suffixes for this filename pattern. -
.<extension>means that there are several (>6) valid extensions for this file type. -
[.gz]means that both the unzipped and gzipped versions of the extension are valid.
If collected, blood measurements of radioactivity are be stored in
Tabular files and located in
the pet/ directory along with the corresponding PET data.
The REQUIRED recording entity is used to distinguish sampling methods.
For example, if an autosampler is used to record continuous blood samples,
and manual measurements are also taken,
then the files may have recording labels autosampler and manual,
respectively.
If the sampling method is unknown, then recording SHOULD be set as manual.
All blood measurements should be reported according to a single time-scale
in relation to time zero defined by the PET data (Figure 1).
All definitions used below are in accordance with
Innis et al. 2007 (doi:10.1038/sj.jcbfm.9600493).
Some metadata about the recording MUST be provided in an additional JSON file.
| Key name | Requirement Level | Data type | Description |
|---|---|---|---|
| PlasmaAvail | REQUIRED | boolean | Boolean that specifies if plasma measurements are available. Must be one of: "true", "false". |
| MetaboliteAvail | REQUIRED | boolean | Boolean that specifies if metabolite measurements are available. If true, the metabolite_parent_fraction column MUST be present in the corresponding *_blood.tsv file. Must be one of: "true", "false". |
| WholeBloodAvail | REQUIRED | boolean | Boolean that specifies if whole blood measurements are available. If true, the whole_blood_radioactivity column MUST be present in the corresponding *_blood.tsv file. Must be one of: "true", "false". |
| DispersionCorrected | REQUIRED | boolean | Boolean flag specifying whether the blood data have been dispersion-corrected. NOTE: not customary for manual samples, and hence should be set to false. Must be one of: "true", "false". |
| WithdrawalRate | RECOMMENDED | number | The rate at which the blood was withdrawn from the subject. The unit of the specified withdrawal rate should be in "mL/s". |
| TubingType | RECOMMENDED | string | Description of the type of tubing used, ideally including the material and (internal) diameter. |
| TubingLength | RECOMMENDED | number | The length of the blood tubing, from the subject to the detector in meters. |
| DispersionConstant | RECOMMENDED | number | External dispersion time constant resulting from tubing in default unit seconds. |
| Haematocrit | RECOMMENDED | number | Measured haematocrit, meaning the volume of erythrocytes divided by the volume of whole blood. |
| BloodDensity | RECOMMENDED | number | Measured blood density. Unit of blood density should be in "g/mL". |
The following metadata SHOULD or MUST be provided if corresponding flags are true.
| Key name | Requirement Level | Data type | Description |
|---|---|---|---|
| PlasmaFreeFraction | RECOMMENDED if PlasmaAvail is true |
number | Measured free fraction in plasma, meaning the concentration of free compound in plasma divided by total concentration of compound in plasma (Units: 0-100%). Must be a number greater than or equal to 0 and less than or equal to 100. |
| PlasmaFreeFractionMethod | RECOMMENDED if PlasmaAvail is true |
string | Method used to estimate free fraction. |
| MetaboliteMethod | REQUIRED if MetaboliteAvail is true |
string | Method used to measure metabolites. |
| MetaboliteRecoveryCorrectionApplied | REQUIRED if MetaboliteAvail is true |
boolean | Metabolite recovery correction from the HPLC, for tracers where it changes with time postinjection. If true, the hplc_recovery_fractions column MUST be present in the corresponding *_blood.tsv file. Must be one of: "true", "false". |
The following columns are defined for _blood.tsv files.
The time column MUST always be the first column.
| Column name | Requirement Level | Data type | Description |
|---|---|---|---|
| time | REQUIRED | number | Time, in seconds, relative to TimeZero defined by the *_pet.json. For example, 5. |
| plasma_radioactivity | OPTIONAL, but REQUIRED if PlasmaAvail is true |
number | Radioactivity in plasma, in unit of plasma radioactivity (for example, kBq/mL). |
| metabolite_parent_fraction | OPTIONAL, but REQUIRED if MetaboliteAvail is true |
number | Parent fraction of the radiotracer (0-1). Must be a number greater than or equal to 0 and less than or equal to 1. |
| metabolite_polar_fraction | OPTIONAL, but RECOMMENDED if MetaboliteAvail is true |
number | Polar metabolite fraction of the radiotracer (0-1). Must be a number greater than or equal to 0 and less than or equal to 1. |
| hplc_recovery_fractions | OPTIONAL, but REQUIRED if MetaboliteRecoveryCorrectionApplied is true |
number | HPLC recovery fractions (the fraction of activity that gets loaded onto the HPLC). |
| whole_blood_radioactivity | OPTIONAL, but REQUIRED if WholeBloodAvail is true |
number | Radioactivity in whole blood samples, in unit of radioactivity measurements in whole blood samples (for example, kBq/mL). |
| Additional Columns | NOT ALLOWED | n/a |
Additional columns are not allowed. |
As with all tabular files,
additional columns MAY be defined in _blood.json.
For clarity, it is RECOMMENDED to include the above column definitions in _blood.json,
as shown in the following example.
Example blood data
*_recording-manual_blood.json:
{
"PlasmaAvail": true,
"WholeBloodAvail": true,
"MetaboliteAvail": true,
"MetaboliteMethod": "HPLC",
"MetaboliteRecoveryCorrectionApplied": false,
"DispersionCorrected": false,
"time": {
"Description": "Time in relation to time zero defined by the _pet.json",
"Units": "s"
},
"plasma_radioactivity": {
"Description": "Radioactivity in plasma samples. Measured using COBRA counter.",
"Units": "kBq/mL"
},
"whole_blood_radioactivity": {
"Description": "Radioactivity in whole blood samples. Measured using COBRA counter.",
"Units": "kBq/mL"
},
"metabolite_parent_fraction": {
"Description": "Parent fraction of the radiotracer.",
"Units": "arbitrary"
},
"metabolite_polar_fraction": {
"Description": "Polar metabolite fraction of the radiotracer.",
"Units": "arbitrary"
},
"metabolite_lipophilic_fraction": {
"Description": "Lipophilic metabolite fraction of the radiotracer.",
"Units": "arbitrary"
}
}
*_recording-manual_blood.tsv:
time plasma_radioactivity whole_blood_radioactivity metabolite_parent_fraction metabolite_polar_fraction
0 0 0 1 0
145 43.31 33.79 0.5749 0.1336
292 48.96 37.42 0.3149 0.2746
602 39.84 32.05 0.1469 0.3548
1248 37.38 31.52 0.073 0.444
1785 36.40 28.83 0.078 0.429
2390 33.13 26.32 0.061 0.453
3059 30.83 25.22 0.049 0.473
4196 27.28 21.98 0.036 0.503
5407 22.70 19.49 0.032 0.523
7193 19.71 15.70 0.02 0.559